
by Teichlab
Wellcome Sanger Institute
Wellcome Sanger Institute
Part of Human Cell Atlas

| Summary | Interactive Viewer | Norm H5AD Data | Raw H5AD Data | RNA Velocyto |
|---|---|---|---|---|
| Full single cell RNA-seq dataset of 428K intestinal cells from fetal, pediatric, adult donors, and up to 11 intestinal regions. Click here to access the paper. | pageview | file_download | file_download | file_download |
| Explore lineages below: | ||||
| Epithelium | pageview | file_download | file_download | |
| Mesenchyme | pageview | file_download | file_download | |
| Enteric neurons | pageview | file_download | file_download | |
| Endothelium | pageview | file_download | file_download | |
| T/NK cells | pageview | file_download | file_download | |
| B cells | pageview | file_download | file_download | |
| Myeloid | pageview | file_download | file_download | |
| Summary | Interactive Viewer | Norm H5AD Data | Raw H5AD Data |
|---|---|---|---|
| Single-cell transcriptomes for 62,849 cells isolated from 6-11 weeks post-conception developing human gut. This data includes intestinal cells from duo-jejunum, ileum and colon. | pageview | file_download | file_download |
| Single-cell transcriptomes isolated from terminal ileum of childhood onset Crohn’s disease and matched healthy controls. This map includes total of 22,500 cells from 8 control samples and 7 Chron’s disease samples. | pageview | file_download | file_download |
 Human enteric nervous system development. Colonisation of the myenteric plexus by enteric neural progenitors.
pageview
Human enteric nervous system development. Colonisation of the myenteric plexus by enteric neural progenitors.
pageview
| Histology & RNAscope imaging by the Cellular Generation and Phenotyping (CGaP) core facility and Bayraktar Lab. |  |
| Summary | Interactive Viewer | H5AD Data | MTX Counts | CSV Gene Names | CSV Metadata |
|---|---|---|---|---|---|
The colon, as a barrier tissue, represents a unique immune environment where immune cells display tolerance toward adiverse community of microbes, which are collectively known as the microbiome.
The published study revealed that not only were there differences between the immune cells in different parts of the colon, but that the microbiome also subtly changed, with a broader range of bacteria further down the colon. 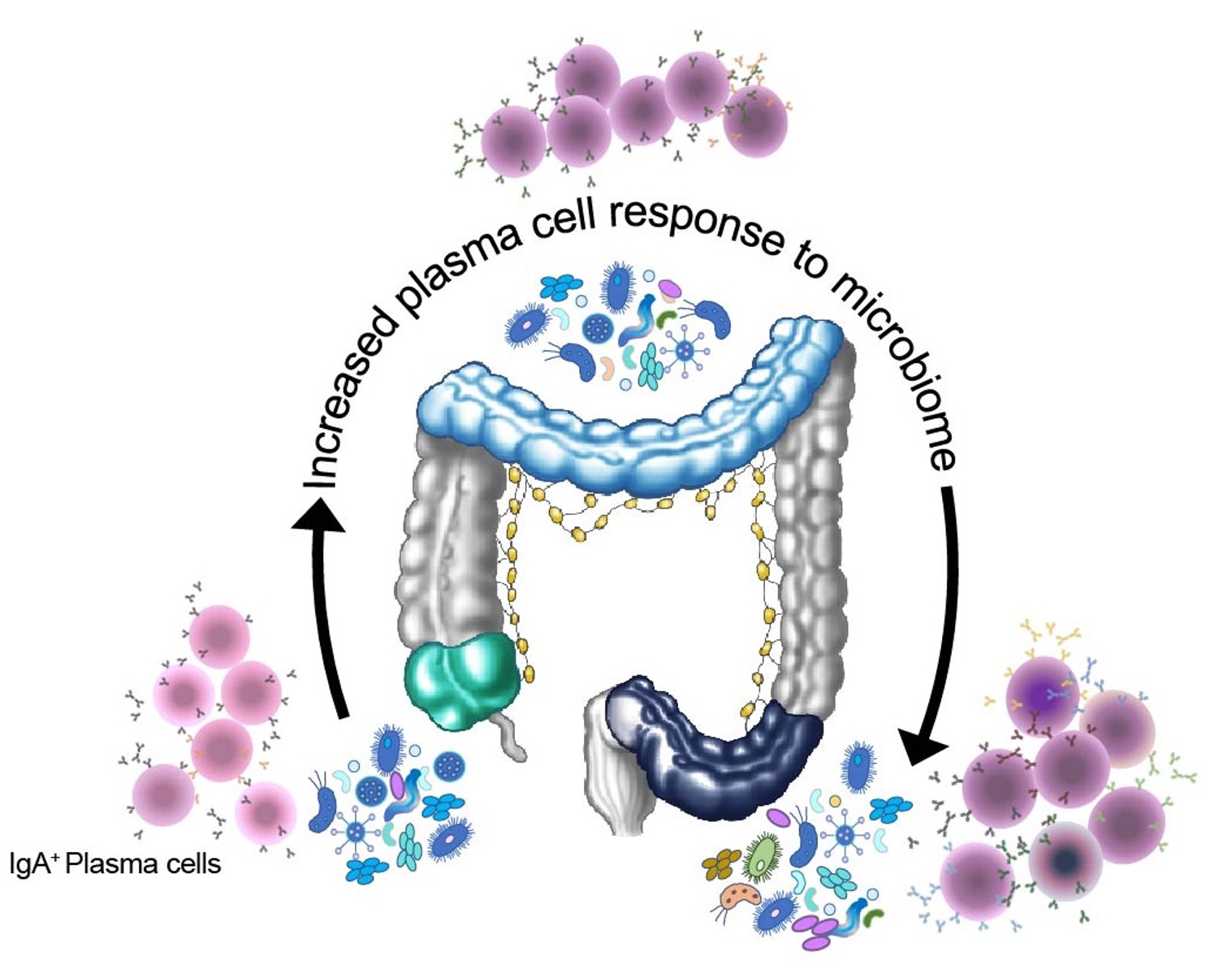
Image Description: Single cell transcriptomes for 41,650 cells isolated from the caeacum, transverse colon and sigmoid colon of 5 individuals. Single cell transcriptomes for 41,650 cells isolated from the caeacum, transverse colon and sigmoid colon of 5 individuals. |
pageview | file_download | file_download | file_download | file_download |